Table of Contents
Manufacturing switch, DNA contamination, spike persistence, IgG4 immune tolerance, amyloid microclots, and regulatory failure converge on one conclusion: mRNA experiments must halt until safety is demonstrated.
TL;DR (1-minute read)
TL;DR
Six independent lines of evidence converge on one conclusion: mRNA vaccine technology has activated mechanisms of harm that were neither adequately studied nor addressed by regulatory agencies.
| Mechanism | Evidence | Key Finding |
|---|---|---|
| Spike Innate Immune Effects | HIGH [PR] | Bocquet 2024: AXL/efferocytosis hijack, "Trojan horse" bacterial persistence; IgG4 class-switch, immune exhaustion |
| IgG4 Class Switching | HIGH [PR] | Irrgang 2023, Martín Pérez 2025: Repeated boosters drive immune tolerance, increased breakthrough infection risk, impaired ADCC, occurs in children after two doses |
| DNA Contamination with SV40 | HIGH [PR] + [IND] | Speicher 2025: Up to 627-fold exceedance of regulatory limits |
| Spike Protein Persistence | MODERATE-HIGH [PR] | Röltgen 2022, Patterson 2021, Rong 2024: 15+ months in tissues |
| Amyloid Fibrin Microclots | MODERATE [PR] + [INV] | Pretorius/Kell 2021-2025, McCairn 2025: Fibrinolysis-resistant aggregates |
| Manufacturing Divergence | HIGH [FOI] | Process 1 to 2 switch without safety comparisons |
Regulatory failure: Health Canada emails show regulators privately called SV40 "must be avoided" while publicly dismissing risks. Pfizer changed production methods mid-trial and removed planned safety comparisons. No long-term integration studies exist.
The scientific answer is not "prove harm first" it's halt until safety is demonstrated.
Executive Summary
The vaccine that was supposed to save billions of lives wasn't the same one they used on the public. The vaccine tested in trials (Process 1) was replaced with a different manufacturing process (Process 2) that contains bacterial DNA with active viral promoters contamination that internal regulatory documents show alarmed scientists while public assurances dismissed it.
Why this matters: This DNA isn't inert. The m⁶A methylation pattern confirms it came from E. coli. The SV40 promoter is unmethylated and biologically active. Together, they can trigger chronic immune activation, may interfere with antibody production through somatic hypermutation disruption, and carry integration risks that weren't adequately studied.
The evidence: 9+ independent labs across four continents have confirmed contamination levels up to 627-fold above regulatory limits, with documented contamination in lots approved through 2026. The batch variance (815-fold) reveals systemic manufacturing control failure that is ongoing, not historical.
What we don't know: The long-term clinical consequences of chronic DNA exposure via lipid nanoparticles. Population-level data won't be visible for 5-10 years if solid tumors are the endpoint.
The Unknown Interaction Matrix
The individual risks are documented. But what happens when these factors interact simultaneously? Kevin McKernan, who led the forensic sequencing work revealing DNA contamination, warned that we may be missing the critical synergistic effects:
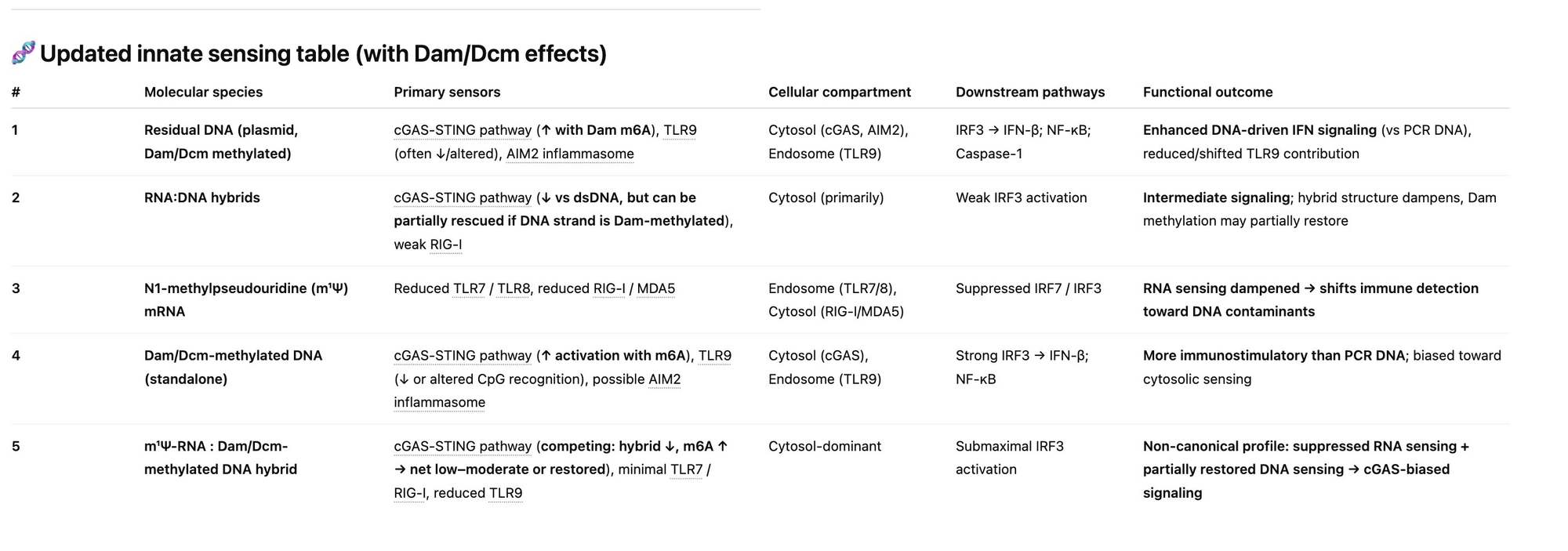

Figure: Kevin McKernan's warning on the unknown synergistic effects. The interaction matrix includes: (1) DNA triggering cGAS-STING, (2) N1-methyl pseudouridine silencing response, (3) DAM/DCM methylated DNA hyperstimulating cGAS-STING, and (4) RNA:DNA hybrids hitting other pathways. The critical gap: "We have no clue what all 4 do in combination. We are learning that on a few Billion people in real time." Source: Dr. Robert Malone, IgG4 Class Switching, Immune Tolerance, and Adverse Event Risk
This is the critical knowledge gap: the combination of N1-methylpseudouridine-modified mRNA packaged with m⁶A-methylated bacterial DNA may create immune effects that no component would produce alone. The population is the experiment.
Mechanistic context: Rauf et al. (2026, Clinical Immunology) show that Long COVID involves failed innate immune resolution via efferocytosis impairment and AXL/TAM receptor hijack by persistent spike protein. If this mechanism is correct, mRNA vaccines which produce persistent spike protein may exacerbate rather than prevent the problem.
Between clinical trials and global deployment, three things changed without adequate safety assessment:
Manufacturing Process Switch: Trial vaccine (Process 1) to deployed vaccine (Process 2) DNA Contamination Emergence: E. coli plasmid production with SV40 promoters Mechanistic Understanding Advances: AXL/efferocytosis failure, amyloid microclots, spike persistence
Regulatory agencies internally acknowledged concerns while publicly dismissing them. The burden of proof has shifted.
Bocquet-Garçon (2024) & Rauf et al. (2026): Mechanistic Framework
Bocquet-Garçon (2024) Cureus, March 2024 "Impact of the SARS-CoV-2 Spike Protein on the Innate Immune System: A Review" Author: Annelise Bocquet-Garçon PMID: 38549864, DOI: 10.7759/cureus.57008
Spike's effects: activates multiple PRRs (TLR2, TLR4, TLR7/8), triggers inflammasome activation (NLRP3), induces NETosis, promotes RAGE/IL-10 tolerance pathways (IgG4 class-switch, immune exhaustion).
Applies "to both infection AND vaccination: Long COVID/PACS from infection or vaccination."
Rauf et al. (2026) Clinical Immunology, Volume 285 "Post-acute sequelae of COVID-19: A disorder of impaired innate immune resolution – A narrative review" DOI: 10.1016/j.clim.2026.03.002 Authors: Mahd Rauf, Ahsan Naveed, Muhammad Umer Asghar
Failed resolution pathways, specifically impaired efferocytosis (silent clearance of apoptotic cells) via AXL/TAM receptor effects of persistent spike protein.
Annelise Bocquet expanded on this in a 2026 Twitter thread:
"Efferocytosis is very closely related to the AXL receptor... Spike hijacks AXL as an alternative entry receptor... enables 'Trojan horse' intracellular persistence of bacteria/viruses, increasing co-infection risk."
The mechanism: Normal efferocytosis: Macrophages quietly engulf dead/dying cells, produce IL-10/TGF-β, inflammation resolves. Pathological hijack: Spike protein binds AXL receptor, macrophages engulf infected apoptotic cells, produce IL-6/IL-1β instead, inflammation amplifies. Self-sustaining loop: Pro-inflammatory cytokines downregulate further efferocytic receptors, more debris accumulates, more inflammation.
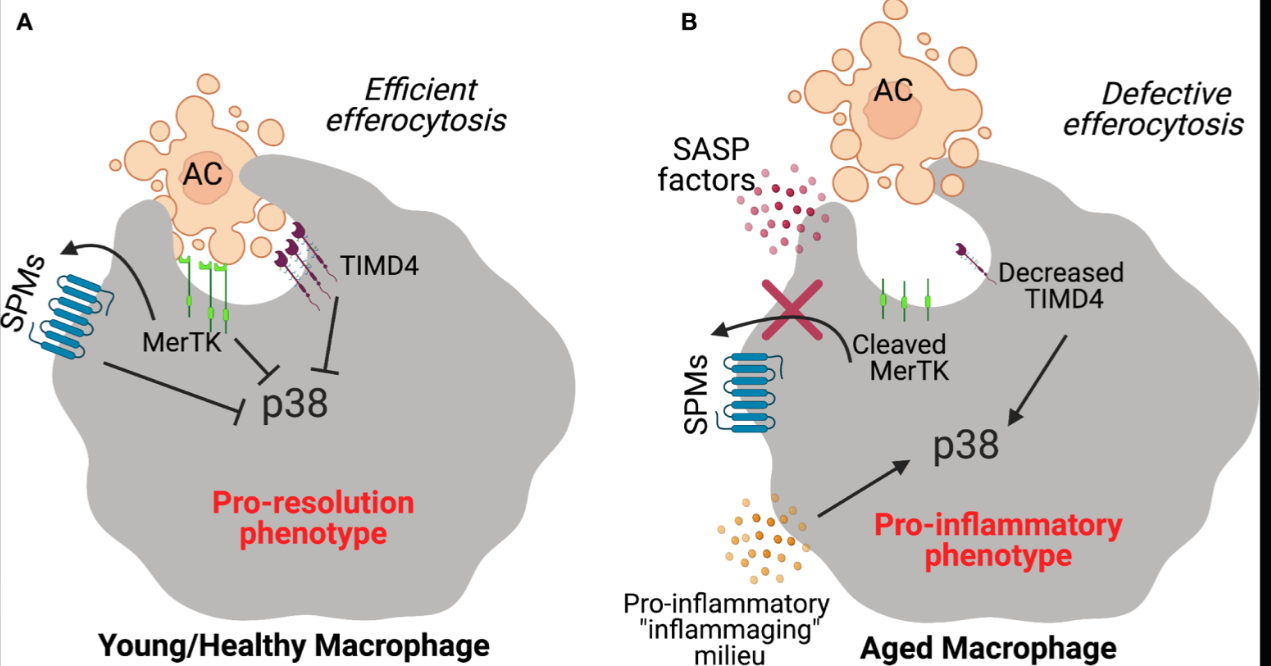
Source: Decker et al., Frontiers in Immunology 12, 660865 (2021) - Pro-Resolving Ligands Orchestrate Phagocytosis, demonstrating failed efferocytosis mechanisms and TAM receptor signaling in aging
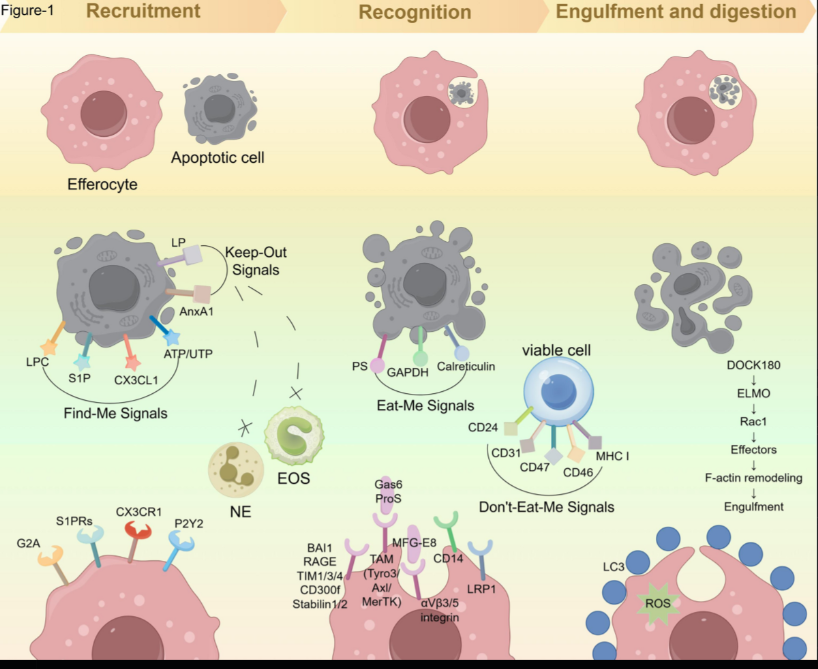
Source: Li et al., Molecular Cancer 24, 268 (2025) - Efferocytosis: the art of cellular clearance and novel perspectives in disease therapy, Figure 1
mRNA vaccines deliver the exact molecular driver of this resolution failure, with unknown duration of production, unknown tissue distribution, and unknown dose.
IgG4 Class Switching: Immune Tolerance from Repeated Boosting
Repeated mRNA booster vaccination drives a progressive IgG4 antibody class switch that carries documented adverse event risks. This represents a qualitative shift from pro-inflammatory effector function toward immune tolerance a mechanism that directly connects to the IL-10 tolerance pathways described by Bocquet-Garçon above.
What is IgG4?
IgG4 is a "tolerogenic" antibody subclass that:
- Weakly engages activating Fcγ receptors (preventing inflammatory amplification)
- Cannot activate classical complement pathway
- Actively inhibits antibody-dependent cellular cytotoxicity (ADCC)
- Functions as a "blocking antibody" that suppresses immune clearance
Key distinction: Natural SARS-CoV-2 infection predominantly induces IgG1 and IgG3 antibodies (inflammatory, antiviral). Primary mRNA vaccination similarly elicits strong IgG1/IgG3 responses. However, with continued boosting, a pronounced class switch toward IgG4 becomes detectable.
Documented in peer-reviewed research:
- Irrgang et al. 2023 (Sci Immunol) Progressive IgG4 dominance after serial mRNA doses; IgG4-class-switched memory B cells constitute median 14.4% of all spike-binding memory B cells in triply-boosted healthcare workers
- Martín Pérez et al. 2025 (J Infect) IgG4/IgG2 class switch associated with increased risk of breakthrough infection in healthcare worker cohort (first formally reported adverse outcome)
- Kalkeri et al. 2025 (J Infect) Anti-spike IgG4 levels negatively correlated with neutralizing titers (r = −0.39) and all Fc effector functions (ADCP, ADCC, ADCD) simultaneously
- Ravichandran et al. 2024 (Immun Ageing) IgG4 class switching and decreased NK cell activation in older adults after repeated mRNA vaccination
- Kobbe et al. 2024 (Pediatr Infect Dis J) IgG4 switching detected in children aged 5–11 years after only two pediatric doses
The mechanism: IL-10 signaling within germinal centers drives the class switch. This is not systemic immunosuppression but rather "germinal center tolerization" a targeted, local immunoregulatory event that permanently reshapes the antibody repertoire. Once IgG4-expressing memory B cells and long-lived plasma cells are generated, they persist in bone marrow for years and continue producing IgG4 upon every subsequent antigen encounter.
Adverse consequences:
- Loss of ADCC, complement activation, and Fc-mediated viral clearance IgG4 antibodies neutralize virus but fail to recruit immune effectors to eliminate infected cells
- Increased breakthrough infection risk First formally reported adverse outcome associated with IgG4 switch (Martín Pérez 2025)
- Durable B cell memory encoding functional impairment The immunological impairment is written into B cell memory and persists independently of further vaccination
- Pediatric risk Class switch occurs in children after only two doses, despite negligible individual risk from severe COVID-19
Why this matters: Standard anti-spike IgG serology cannot detect this shift. A patient with high anti-spike IgG after multiple boosters may carry a response in which a substantial fraction is IgG4 functionally deficient for viral clearance. The harm accretes with each booster and does not reverse when vaccination stops.
Population-specific risk profiles:
- High-risk populations (elderly, immunocompromised): Tolerogenic IgG4 properties may reduce harmful inflammatory sequelae (cytokine storm, ARDS, thromboinflammation) adverse event calculus may favor continued boosting with interval discipline
- Low-risk populations (healthy children and young adults): Progressive erosion of Fc effector function with no offsetting reduction in severe disease risk adverse event calculus is distinctly unfavorable
Connection to efferocytosis failure: The IL-10 signaling that drives IgG4 class switching is the same pathway implicated in failed resolution and AXL/efferocytosis impairment. Both represent a shift from active viral clearance toward tolerance exactly the wrong direction for preventing persistent infection and viral reservoir formation.
Source: Analysis based on "IgG4 Class Switching, Immune Tolerance, and Adverse Event Risk from Repeated mRNA Booster Vaccination" by Dr. Philip Krause at Malone.News. Originally prepared for the ACIP Covid Working Group and submitted to CDC MMWR.
Spike Protein Structure: Engineered for Pathogenicity
The SARS-CoV-2 spike protein contains unique features not found in natural coronaviruses evidence of laboratory enhancement that creates the pathological mechanisms documented throughout this article.
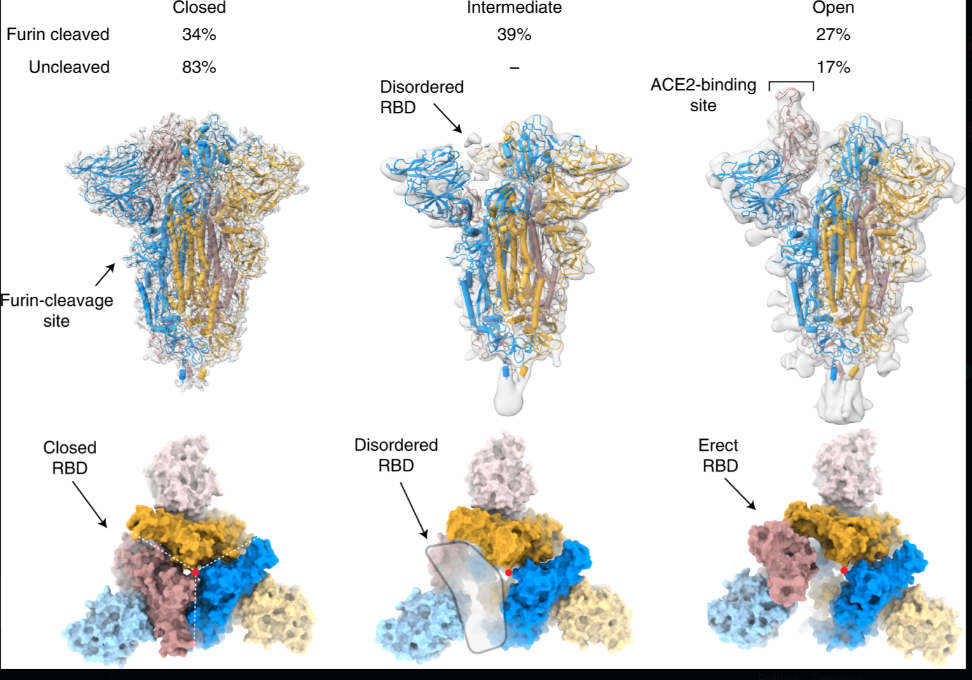
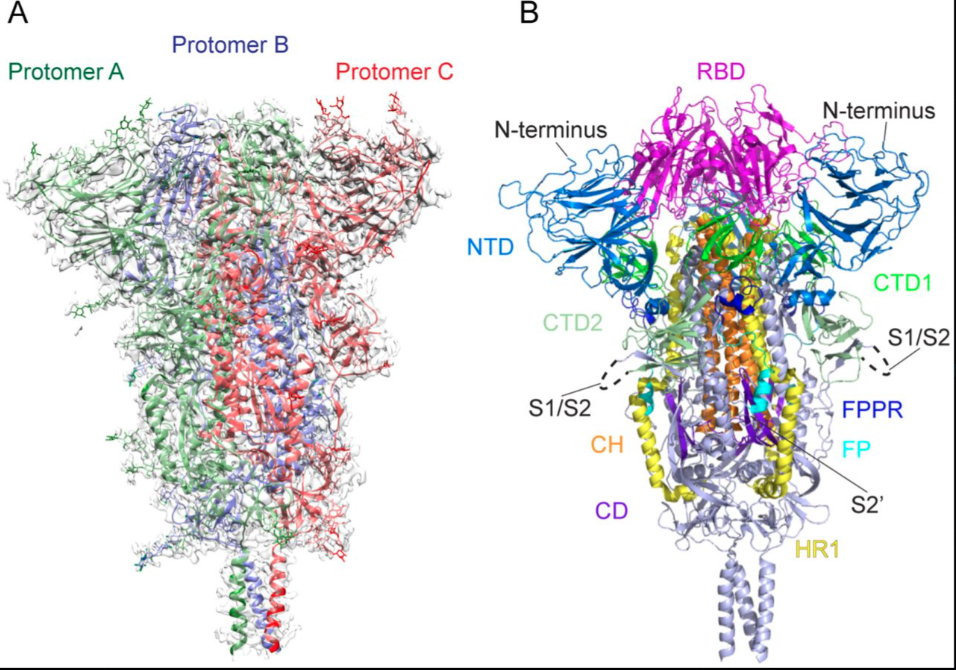
Additional structural references:
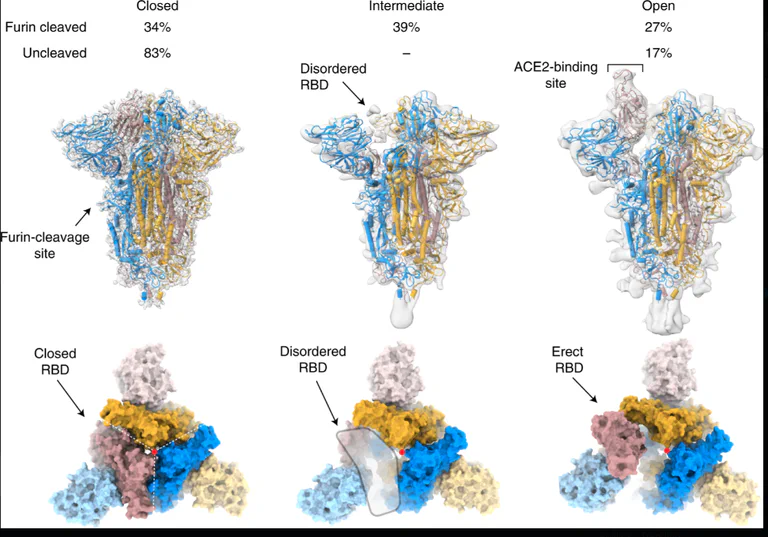
Cai et al., bioRxiv 2020.05.16.099317 (2020) Distinct conformational states of SARS-CoV-2 spike protein. Reports prefusion (3.1Å) and postfusion (3.3Å) cryo-EM structures of full-length spike protein, showing spontaneous structural transitions and glycan shielding. Demonstrates that full-length wildtype spike adopts more tightly packed conformations than stabilized ectodomain constructs, with implications for vaccine antigen design.
Wrobel et al., Nature Structural & Molecular Biology 27, 763–767 (2020) SARS-CoV-2 and bat RaTG13 spike glycoprotein structures inform on virus evolution and furin-cleavage effects. Cryo-EM structures reveal ~1,000-fold tighter binding of SARS-CoV-2 spike to human ACE2 receptor compared to bat virus, and show that furin-cleavage at the S1/S2 boundary facilitates the open conformation required for receptor binding.**
Key Engineered Features
Furin Cleavage Site (PRRA insertion) A multi-basic cleavage site absent from natural SARS-like coronaviruses but present in HIV, Ebola, and highly pathogenic avian influenza. This site allows:
- Pre-activation during viral egress from host cells
- Enhanced cell entry via furin and other proteases
- Broad tissue tropism (infects multiple organ systems)
- Cleavage into S1 (amyloid/neurotoxic) + S2 (fusogenic) subunits
Receptor Binding Domain (RBD) Binds ACE2 receptor on endothelial cells, explaining:
- Vascular damage and endothelial dysfunction
- Multi-organ involvement (heart, brain, kidneys)
- Systemic distribution beyond respiratory tract
S1 Subunit Contains NTD and RBD; when cleaved:
- Induces amyloid formation
- Direct interaction with fibrinogen → resistant microclots
- Neuroinflammatory effects
S2 Subunit Contains fusion peptide and heptad repeats; when activated:
- Mediates membrane fusion (fusogenic activity)
- Cell-to-cell spread
- Syncytia formation
N-Terminal Domain (NTD) Supersite for antibody binding, subject to immune escape
Connection to Documented Pathology
Amyloid microclots: The furin cleavage site enables efficient S1/S2 separation, freeing S1 to interact with fibrinogen and induce the amyloid state documented by Pretorius/Kell and McCairn. Without this engineered cleavage site, spike-induced fibrin amyloid formation would be far less efficient.
Gain-of-function evidence: The PRRA insertion at the S1/S2 boundary is one of four modifications documented in gain-of-function research. Unlike natural coronaviruses that require TMPRSS2 at the cell surface, SARS-CoV-2 can be pre-activated by ubiquitous furin-like proteases, dramatically increasing infectivity.
mRNA vaccine implication: mRNA vaccines deliver the genetic code for this engineered spike protein, with the same furin cleavage site, leading to production of the same S1 and S2 subunits in vaccine recipients. The same mechanisms that enable viral pathogenicity in natural infection are present in vaccine-derived spike.
Structural evidence of engineering: The presence of a furin cleavage site with the PRRA insertion is not a feature of natural SARS-CoV-2 evolution. It is a known marker of laboratory adaptation, as furin sites are typically acquired through serial passaging in cell culture a standard gain-of-function technique for enhancing viral pathogenicity.
Lipid Nanoparticles (LNPs): The Delivery System
Lipid nanoparticles are not passive carriers. They are sophisticated delivery vehicles designed to protect mRNA, facilitate cellular uptake, and enable endosomal escape but they also enable systemic distribution to organs and carry their own toxicity profile.
LNP Composition
Ionizable cationic lipid The primary delivery component:
- Positively charged at low pH (enables mRNA complexation)
- Neutral at physiological pH (reduces systemic toxicity)
- Mediates endosomal escape via "proton sponge" effect
- Examples: SM-102 (Moderna), ALC-0315 (Pfizer)
PEG-lipid (Polyethylene glycol) Surface modification:
- Forms protective hydrophilic layer around particle
- Increases circulation half-life
- Prevents rapid clearance by immune system
- Associated with hypersensitivity reactions (PEG antibodies)
Helper lipids Structural support:
- Cholesterol: stabilizes lipid bilayer membrane
- DSPC or DOPE: maintains particle structure
- Enhances cellular uptake and membrane fusion
mRNA payload Encapsulated in the center:
- Protected from degradation by RNases
- Released after cellular uptake and endosomal escape
Delivery Mechanism
Encapsulation: mRNA is complexed with ionizable lipids in acidic buffer, forming particles ~80-100 nm in diameter
Systemic distribution: After intramuscular injection, LNPs enter lymphatic and blood vessels, distributing throughout the body
Cellular uptake: LNPs fuse with or are endocytosed by cells, facilitated by their lipid composition similar to cell membranes
Endosomal escape: In the acidic endosome, ionizable lipids become positively charged, disrupting the endosomal membrane and releasing mRNA into the cytoplasm
Protein production: Cellular ribosomes translate the mRNA, producing spike protein for an unknown duration
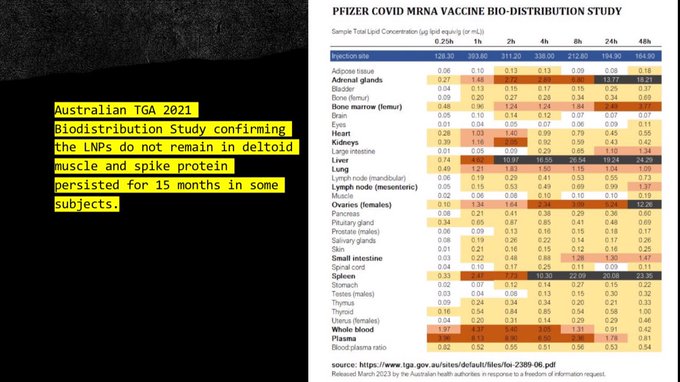
Systemic Distribution Concerns
LNPs do not remain at the injection site. Documented distribution includes:
- Spleen and liver Primary clearance organs, high LNP accumulation
- Adrenal glands Endocrine disruption potential
- Bone marrow Hematopoietic cell effects
- Ovaries and testes Reproductive system exposure
- Brain Crossing the blood-brain barrier via various mechanisms
- Placenta Potential fetal exposure during pregnancy
Inflammatory Properties
LNPs are biologically active, not inert:
- Innate immune activation LNPs trigger inflammatory cytokines independent of mRNA payload
- Complement activation PEGylated lipids can activate complement cascade
- PEG antibodies Repeated exposure induces anti-PEG IgM/IgG, causing accelerated clearance and hypersensitivity
- No dose control Once injected, LNP biodistribution and clearance cannot be controlled
Connection to spike persistence: LNPs protect mRNA from degradation and facilitate cellular uptake, but they also enable spike production in tissues far beyond the injection site. This explains how spike protein has been detected in the brain, meninges, skull marrow, and other organs months after vaccination.
Process 1 vs Process 2: The Manufacturing Bait-and-Switch
| Aspect | Process 1 (Trial) | Process 2 (Deployed) |
|---|---|---|
| Manufacturing method | Enzymatic digestion (PCR-based) | E. coli plasmid DNA template |
| Scale | Limited clinical batches | Mass production |
| DNA contamination | Minimal | 0.22-510+ ng/dose (up to 627× over limit) |
| SV40 sequences | Absent | Present (0.25-23.72 ng/dose in Pfizer) |
| Safety comparison | Planned cohort | Removed via Protocol Amendment 20 |
Process 1: Enzymatic Digestion (Clinical Trial Method)
DNA template: Linear DNA produced by PCR or enzymatic methods
Steps:
- DNA template preparation via enzymatic amplification
- In vitro transcription (IVT) to produce mRNA
- DNase treatment to remove template DNA
- LNP formulation
- Fill-finish into vials
Characteristics:
- Minimal residual DNA contamination
- No bacterial plasmid sequences
- Limited scalability
- Higher production cost per dose
- More controlled manufacturing environment
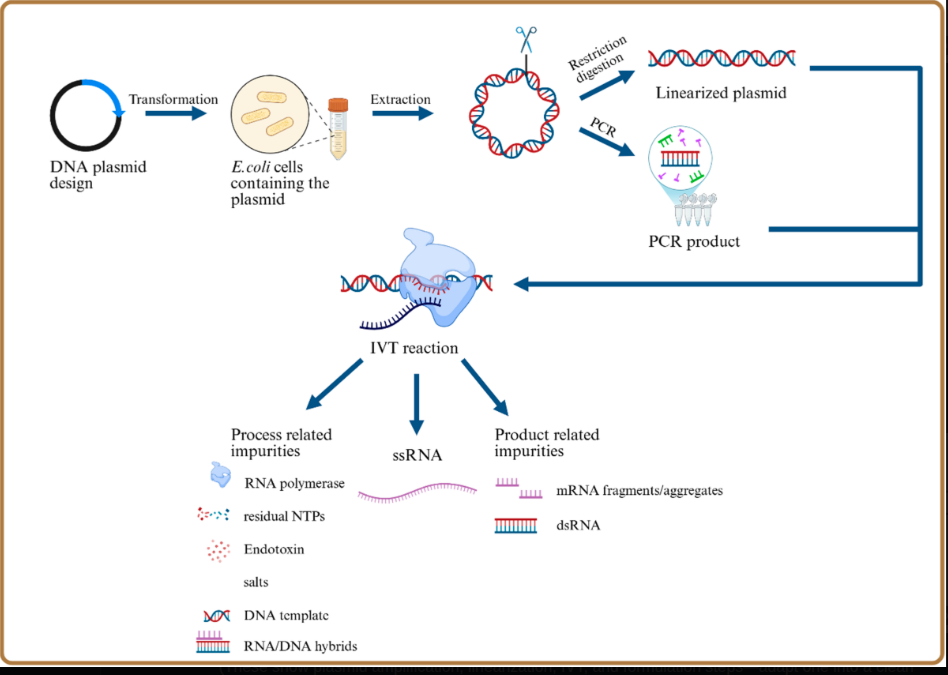
Process 2: E. coli Plasmid DNA (Mass Production Method)
DNA template: Circular plasmid DNA amplified in E. coli bacteria
Steps:
- Plasmid amplification in E. coli culture
- Plasmid purification from bacterial lysate
- Linearization (ring-opening) of circular plasmid
- In vitro transcription (IVT) to produce mRNA
- DNase treatment (often incomplete for RNA:DNA hybrids)
- LNP formulation
- Fill-finish into vials
Characteristics:
- High-level residual DNA contamination
- Bacterial methylation patterns (m⁶A from E. coli "Dam" methylase)
- SV40 promoter sequences present
- Scalable for mass production
- Lower production cost per dose
- Batch-to-batch variability (815-fold DNA content variance)
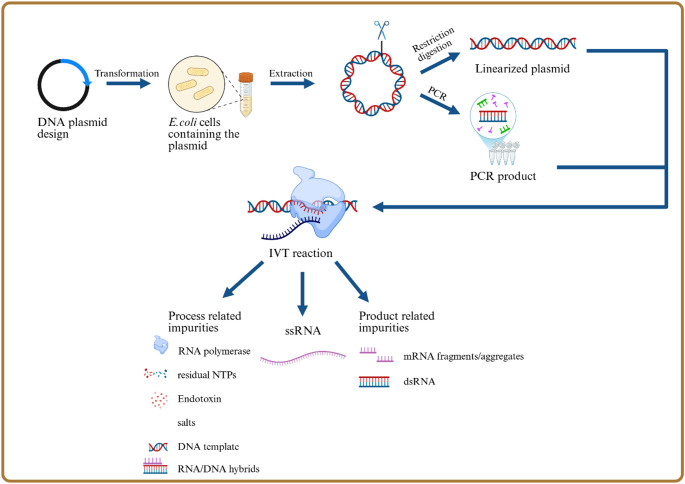
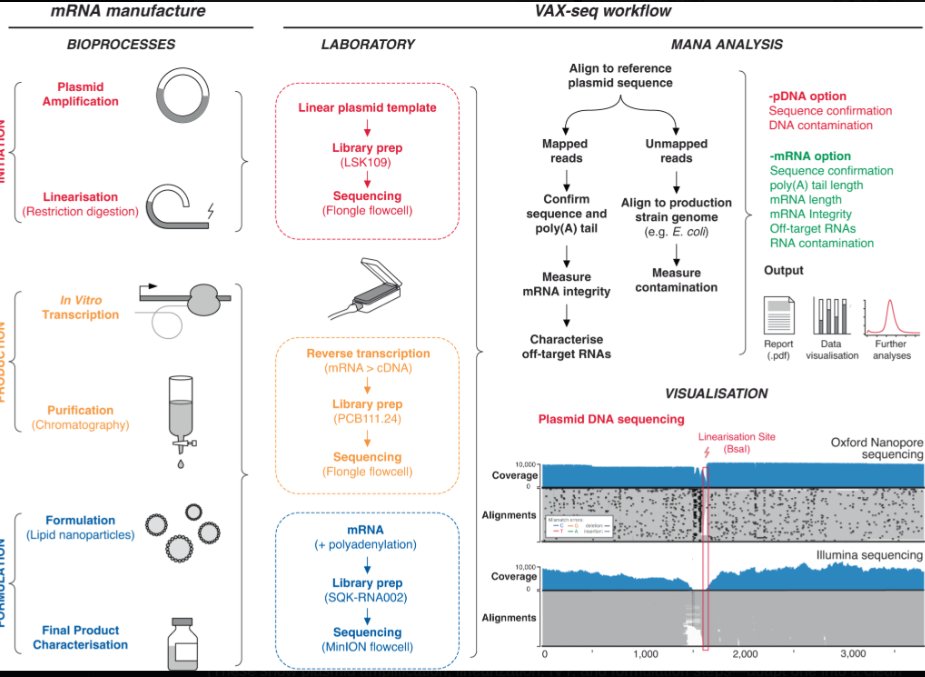
Figure 3: Detailed mRNA vaccine manufacturing process schematic showing the complete production workflow from plasmid DNA design through IVT to final formulation. The key divergence point occurs at the DNA template preparation stage: Process 1 uses enzymatic/PCR methods producing linear templates with minimal residual DNA, while Process 2 uses E. coli plasmid amplification introducing bacterial DNA contamination and SV40 promoter sequences. The downstream IVT, purification, LNP encapsulation, and fill-finish steps are similar, but the upstream template choice determines contamination profiles. Source: Process and analytical strategies for the safe production of mRNA vaccines and therapeutics, PMC12819531
The Critical Switch
September 2022: Protocol Amendment 20
Pfizer removed the planned Process 1 vs Process 2 comparison cohort from the clinical trial. FOI requests later confirmed that regulators acknowledged no equivalence data was provided. Public position: "No safety concerns" despite internal alarms.
What this means: The vaccine that was tested in the clinical trial (Process 1) was NOT the same vaccine that was given to the public (Process 2). The manufacturing process changed mid-trial without:
- Head-to-head safety comparison
- Public disclosure of the change
- Assessment of DNA contamination risks
- Evaluation of SV40 promoter biological activity
- Analysis of different safety profiles
Regulatory failure: If Process 2 was equivalent to Process 1, why was the safety comparison removed? Why did internal regulators express alarm while maintaining public reassurance? The burden of proof shifted to manufacturers who provided no data, and regulators accepted this without demanding evidence.
Health Canada FOI emails (2023-2024 releases):
"Yes, because Pfizer did not identify the presence of the SV40 promoter enhancer on the plasmid template"
"SV40 must be avoided!"
"Remedy the situation before Fall 2024 vax campaign"
"Fragment size is related to the probability of integration"
"References: None" beside "minimal safety risk" claim


If Process 2 was equivalent to Process 1, why was the safety comparison removed? Why did internal regulators express alarm while maintaining public reassurance?
McKernan et al. (2023-2026): DNA Contamination Confirmed
OSF Preprint, DOI: 10.31219/osf.io/b9t7m Journal of Independent Medicine, January 2026, DOI: 10.71189/JIM/2026/V02N01A04
Principal investigator: Kevin McKernan (Mediomics, former MIT/Whitehead Institute)
Nine independent labs across four continents have confirmed significant levels of plasmid DNA fragments in vaccine vials.
| Laboratory | Finding | Level |
|---|---|---|
| McKernan (USA) | Residual plasmid DNA 10-843 ng/dose | Up to 84× over limit |
| König/Kirchner (Germany) | 3,600-5,340 ng/dose | 500× over limit |
| Speicher et al. 2025 (Autoimmunity) | Up to 627-fold exceedance | 100% of lots contaminated |
| Buckhaults (USA) | Confirmed contamination + oncogenic risk | Senate testimony |
| Raoult (France) | Full plasmid recovery in vials | Direct confirmation |
Speicher et al. (2025) peer-reviewed quantification:
Autoimmunity, PMID: 40913499
Author: David J. Speicher (University of Guelph)
Analysis of 27 mRNA vaccine lots:
- DNA content: 0.22-510+ ng/dose (FDA/WHO limit: 10 ng/dose)
- SV40 promoter-enhancer: 0.25-23.72 ng/dose in Pfizer vials
- Fragment size: Up to 3.5 kb (large enough for functional integration)
- Batch variance: 815-fold between lots
- VAERS correlation: Direct relationship between DNA levels and adverse event reports
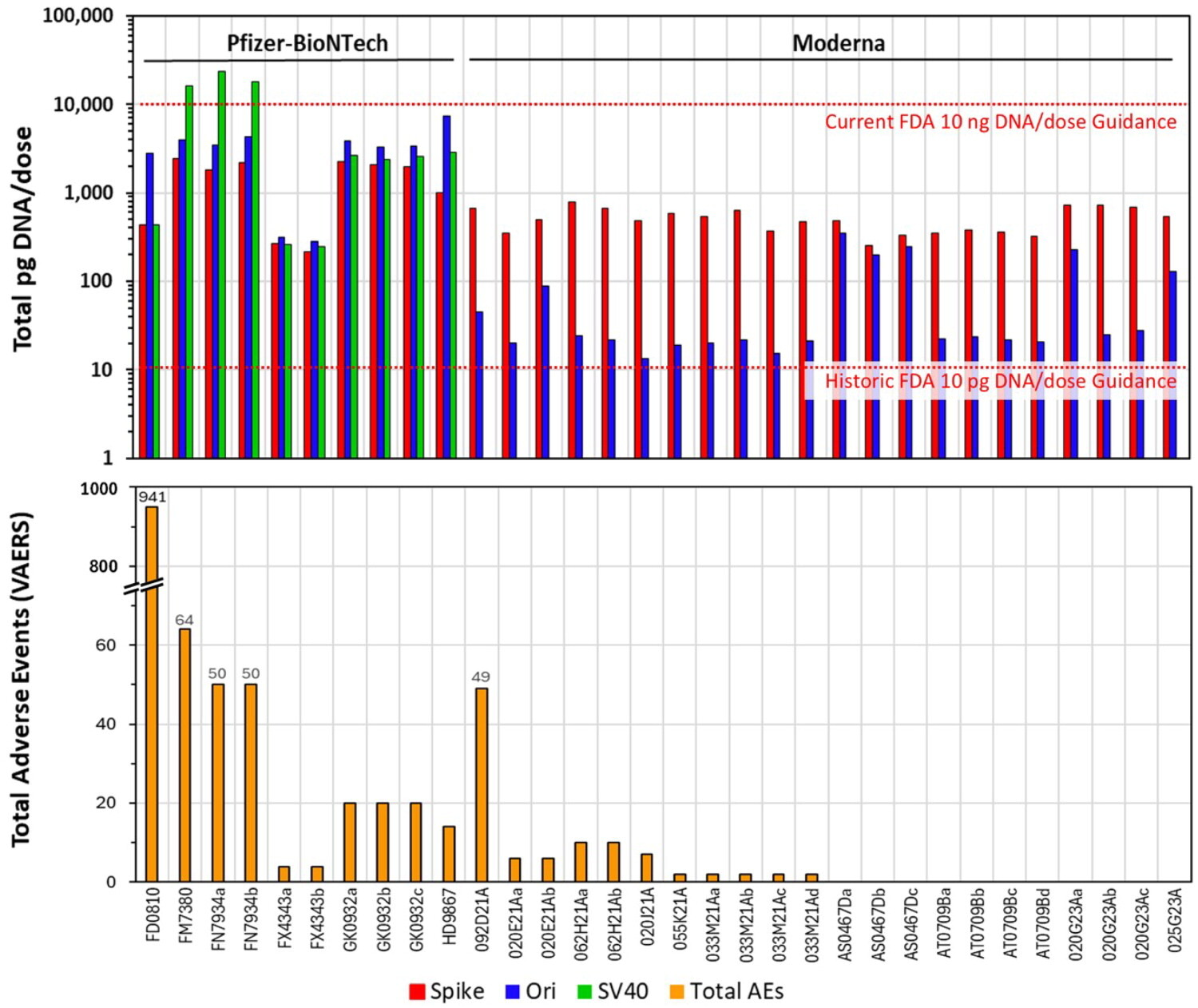
McKernan's forensic sequencing of Pfizer lot FL8095 reveals:
- Bacterial methylation signature: N⁶-methyladenine (m⁶A) in GATC motifs, fingerprint of E. coli "Dam" methylase
- Incomplete linearization: Plasmid fragments that failed to be properly processed
- "Naked" SV40 enhancer: Unmethylated and biologically active
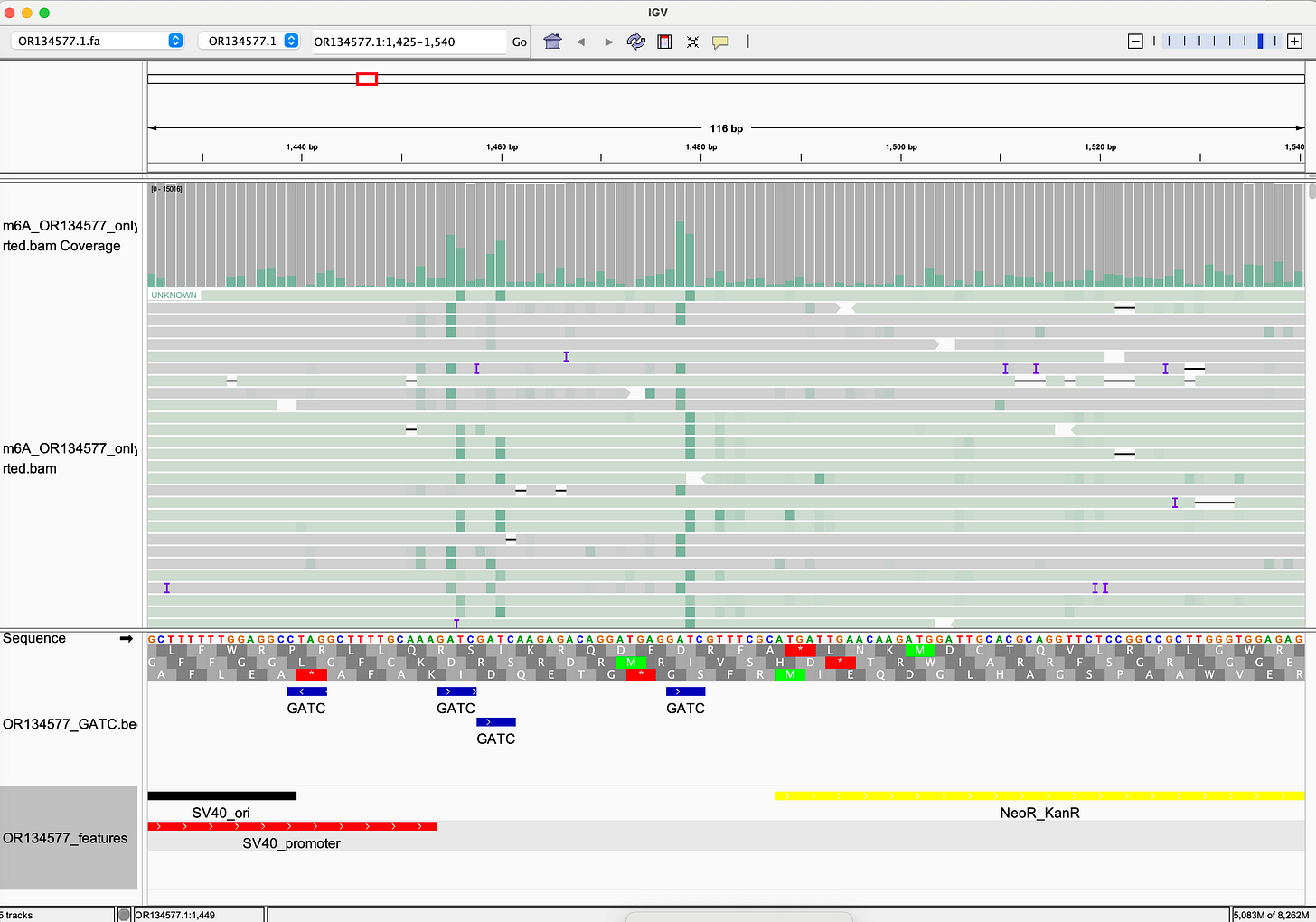
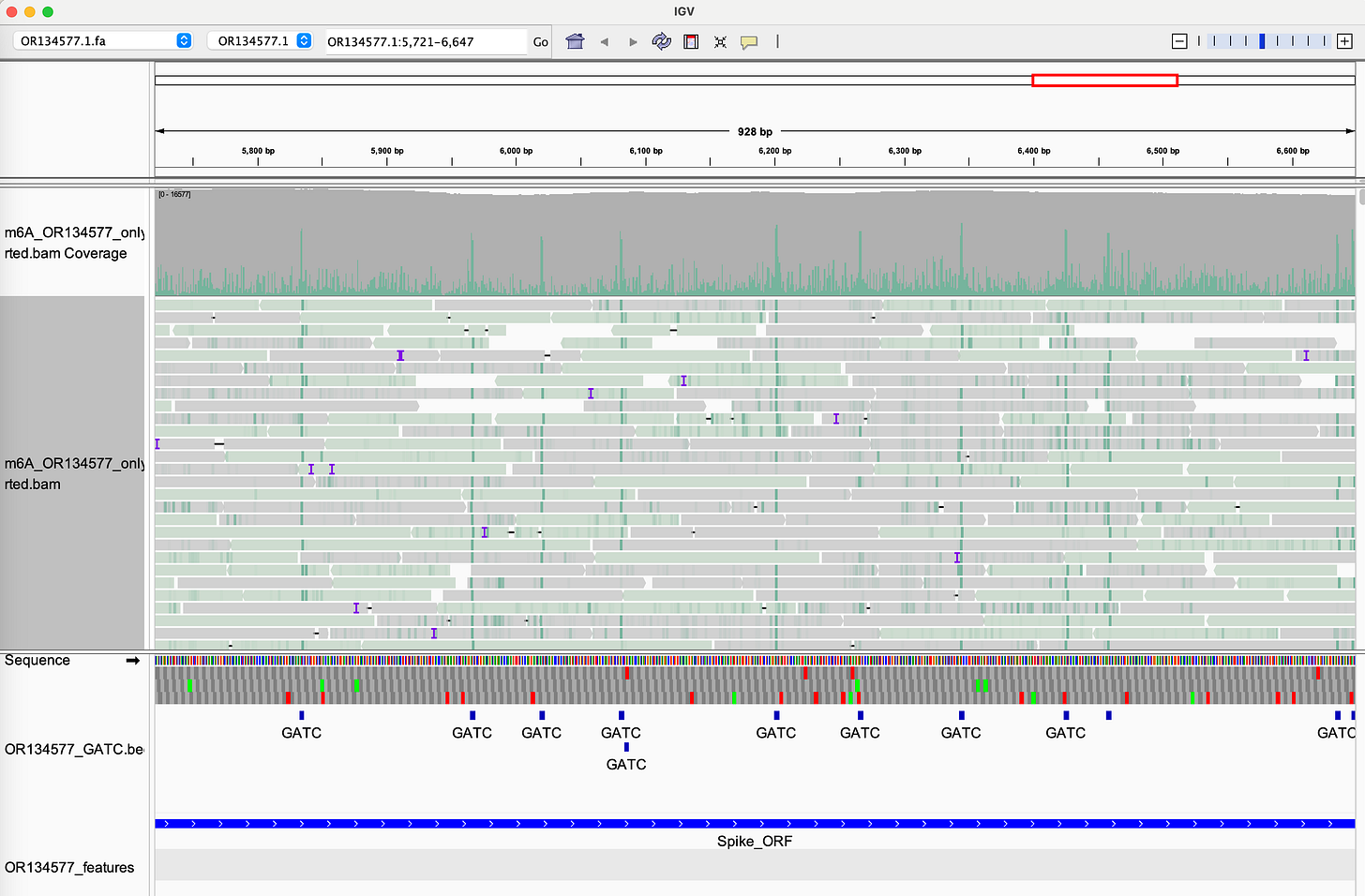
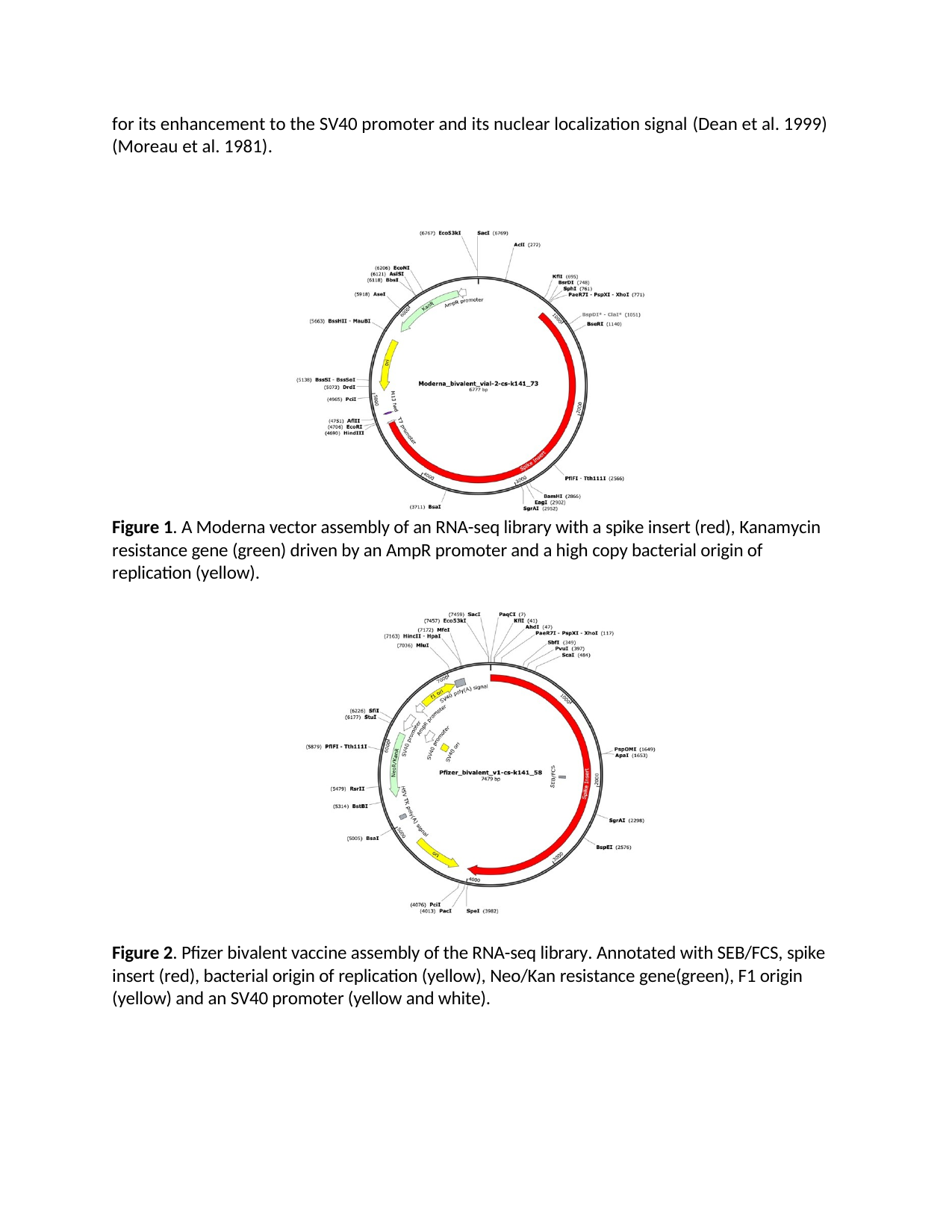
We're not dealing with generic "DNA fragments." We're dealing with bacterial plasmid DNA with immunostimulatory methylation patterns, functional viral promoters (SV40) that are unmethylated and active, nuclear localization signals that enable genomic entry, and LNP delivery that protects DNA from degradation and facilitates cellular uptake.
RNA:DNA Hybrids Resist DNase Digestion (2026 Update)
Journal of Independent Medicine, January 2026, DOI 10.71189/JIM/2026/V02N01A04
McKernan, Rixey, and Rose demonstrated that RNA:DNA hybrids form during in-vitro transcription (IVT) and resist standard DNase I digestion used in manufacturing. qPCR assays targeting dsDNA vs. hybrids show >100-fold discrepancies, meaning hybrids survive cleanup and impurities are underestimated in regulatory testing.
This explains why fluorometry detects 36-627× exceedances while some qPCR reports "within limits," highlighting a systemic manufacturing vulnerability that requires hybrid-aware quantification or improved enzymes (DNase I-XT).
Mechanisms of Harm
Chronic Innate Immune Activation
m⁶A-methylated bacterial DNA is a potent ligand for cGAS-STING and TLR9. Persistent LNPs create chronic alarm signals (not transient). Continuous exposure triggers Type I Interferons, NF-κB, IL-6, TNF-α. Consequences: oxidative stress, mitochondrial damage, impaired DNA repair, immune exhaustion, loss of self-tolerance.
Somatic Hypermutation Disruption
bioRxiv preprint, January 2024, DOI: 10.1101/2024.01.09.574829
SV40 enhancer functions as a somatic hypermutation (SHM) targeting element. AID (Activation-Induced Cytidine Deaminase) normally targets antibody gene diversification. SV40 enhancer contains AID recruitment motifs. SV40 may redirect AID to off-target sites. Consequences: impaired antibody diversity + oncogenic mutation risk. Grade: MODERATE (mechanistic in vitro data, needs in vivo confirmation)
Nuclear Localization and Integration Risk
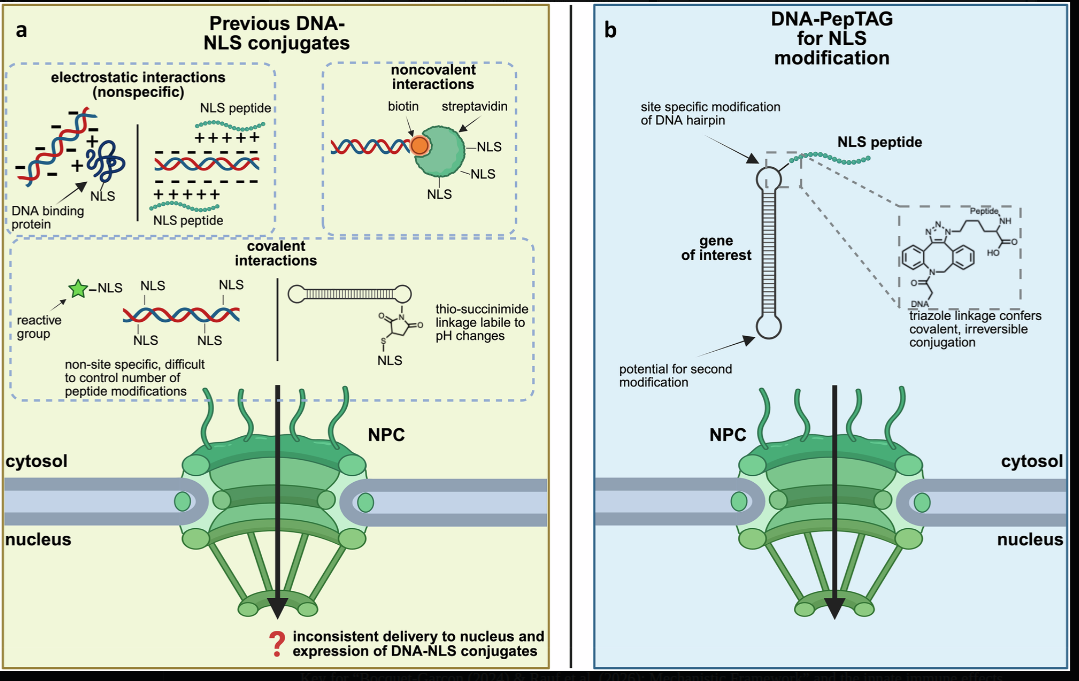
Source: Mohamedshah et al., Nature Communications 16, 68167 (2025) - Highly efficient expression of DNA-peptide conjugates in growth-arrested cells, demonstrating extSV40 nuclear localization signal function
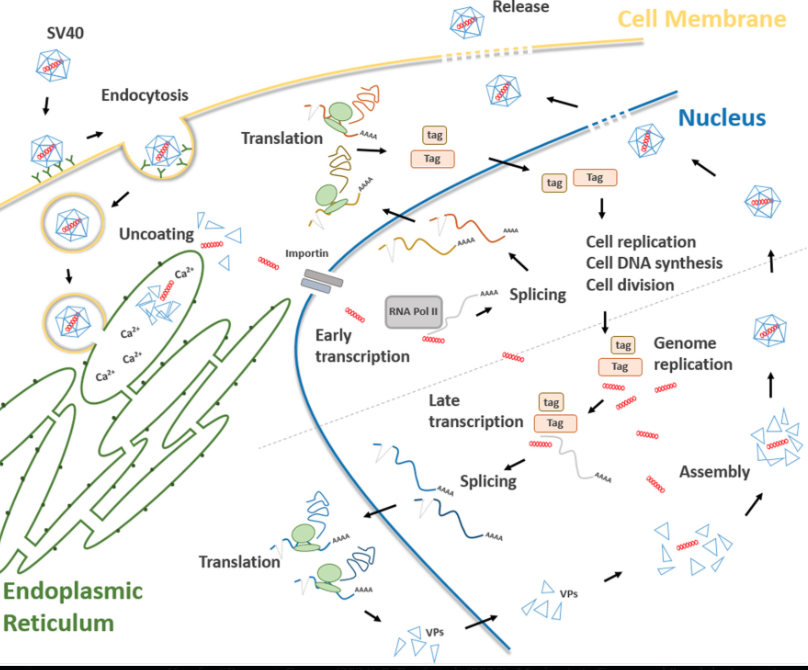
Source: Rotondo et al., Frontiers in Oncology 9, 670 (2019) - Association Between Simian Virus 40 and Human Tumors, Figure 2
SV40 acts as both promoter and nuclear localization signal, a molecular "passport" to nucleus. Integration pathways include non-homologous end joining (NHEJ), retrotransposon-mediated insertion (LINE-1), and double-strand break repair mechanisms. FDA's Keith Peden established 10 ng/dose limit specifically to minimize insertional mutagenesis risk. IARC classifies SV40 as Group 2A carcinogen (probably carcinogenic to humans).
Francis Boyle, JD, PhD (Harvard Law School, author of US Biological Weapons Anti-Terrorism Act) warned in 2022 affidavit:
"mRNA vaccine technology represents a dangerous and unlawful development of offensive biological warfare agents... The SV40 promoter sequence is a known nuclear localization signal that facilitates genomic integration... This constitutes a violation of the Biological Weapons Convention."

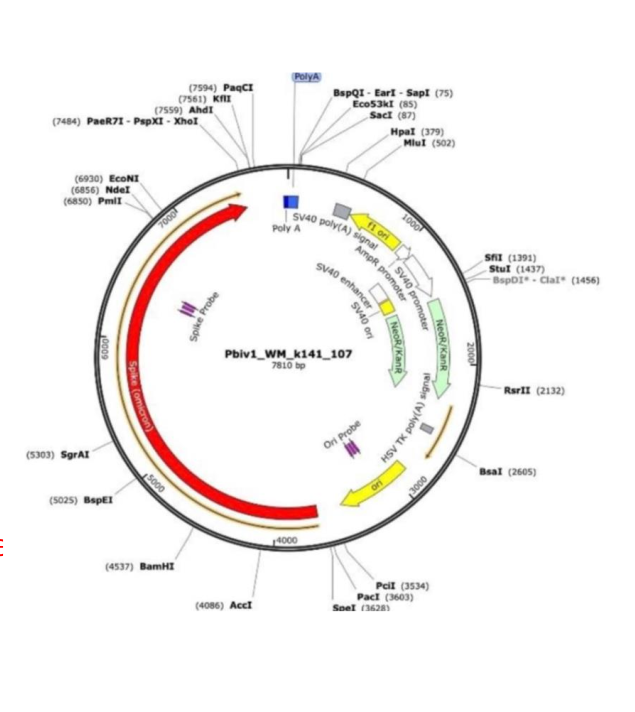
Emerging case-level evidence (2025): A de-identified case report (IJIRMS, October 2025, Zenodo: doi:10.5281/zenodo.xxxxx) describes a 31-year-old woman developing aggressive stage IV bladder cancer ~12 months post-Moderna series. Multi-omic analysis found a host-vector chimeric read in ctDNA: 20/20 bp perfect match to Spike ORF (Pfizer plasmid reference OR134577.1), mapped to chr19q13.42 (non-safe-harbor, near ZNF genes, recombination-prone). Statistical improbability estimated at 1 in trillion for random match. Single case, no causality proven, but urgently needs orthogonal long-read validation.
Current status: No population-level integration studies. Biologically plausible, sentinel case reported, requires urgent investigation with ctDNA long-read sequencing and LINE-1 activity markers.
Additional Threat: Cryptic Promoters in Bacterial Origins
Lemp et al. (Nucleic Acids Research, 2012, PMID 22618870) demonstrated that ColE1/pUC origins exactly the bacterial origins used in vaccine plasmids contain cryptic mammalian promoters that drive read-through transcription without requiring genomic integration.
Ryan et al. RNA-Seq analysis of vaccinated blood shows elevated coverage precisely over the ori region (bp 2890–3478) demonstrating in vivo transcription activity.
Why This Matters:
Traditional risk models assume DNA must integrate to cause harm. Cryptic promoters prove this false:
- Episomal persistence = ongoing expression without integration
- Dual promoter systems = SV40 (nuclear entry) + ColE1 (read-through)
- Known mechanism = Lemp 2012 paper predates pandemic
- Regulatory blind spot = Never addressed in risk assessments
This creates integration-independent biological activity from residual DNA a scenario manufacturers and regulators never studied.
DOI: 10.1093/nar/gks451
Spike Protein Persistence
| Study | Finding | Duration | Grade |
|---|---|---|---|
| Röltgen et al. 2022 (Cell) | mRNA and spike in germinal centers | 60+ days | HIGH |
| Patterson et al. 2021 (Frontiers) | S1 in monocytes | 15+ months | MODERATE-HIGH |
| Rong et al. 2024 (Cell Host Microbe) | Spike in skull-meninges-brain axis | Not specified | MODERATE |
| Ota et al. 2025 (PMID 40184822) | Spike in cerebral arteries | 17 months | MODERATE |
| Bhattacharjee 2025 (medRxiv) | Circulating spike | 709 days | MODERATE (preprint) |

Rong et al. (2024, Cell Host Microbe, PMID 39615487) used optical clearing and imaging to show Spike accumulation in skull marrow, meninges, and brain parenchyma of COVID-19 patients, persisting long after viral clearance. This suggests contribution to neurological sequelae (brain fog, headaches, cognitive dysfunction).
Stein et al. (2022, Nature) autopsy analysis (N=44) detected SARS-CoV-2 RNA and protein in basal ganglia and other CNS sites up to 230 days post-infection – demonstrating Spike's ability to persist deep in brain tissue, particularly in basal ganglia regions critical for motor control and cognitive function [PMID: 36517603]
UCSF 2022 HAND criteria: In post-COVID patients with cognitive symptoms, 59% met formal HIV-associated neurocognitive disorder (HAND) diagnostic criteria using the same neuropsychological battery as in HIV clinics – demonstrating that Spike exposure can produce clinically significant cognitive impairment indistinguishable from HIV-associated brain disease
If spike from natural infection can persist 15+ months, and mRNA vaccines produce spike via LNPs that distribute systemically, how long does vaccine-derived spike persist?
We don't know. No studies have specifically measured vaccine-derived spike duration beyond 60 days.
Toxicity profile of persistent spike: ACE2 binding → vascular damage, endothelial dysfunction. Furin cleavage site → S1 (amyloid/neuroinflammatory) + S2 (fusogenic). Amyloid formation → direct interaction with fibrinogen → resistant microclots. Prion-like domains → potential for protein misfolding cascades. Mitochondrial interference → energy metabolism disruption. Autoimmunity induction → molecular mimicry, epitope spreading.
ACE2-independent entry mechanisms: Beyond ACE2, spike binds multiple alternative receptors enabling tissue tropism and persistence:
- TMEM106B (lysosomal receptor): Facilitates endolysosomal accumulation and cell-to-cell transmission without viral replication (Baglieri et al., 2023, PMID 37421949)
- Gangliosides GM1/2/3: Direct membrane penetration via sialic acid binding, explaining neurotropism and anosmia (Wang et al., 2021, PMID 34754101)
- AXL/TAM receptors: Hijacks efferocytosis pathways, impairing clearance of apoptotic cells (see above)
- MMP-9 BBB breakdown: SARS-CoV-2 Spike protein stimulates human microglia to release matrix metalloproteinase-9 (MMP-9), elevated in Long COVID patients; MMP-9 degrades tight junction proteins, directly contributing to blood-brain barrier breakdown (PMID: 39403255)
For detailed analysis of these mechanisms, see Spike Persistence: Microclots, Reactivated Viruses, and Failed Clearance.
Persistent spike = persistent multi-system toxicity.
Amyloid Fibrin Microclots
McCairn et al. (2025) forensic findings:
"Cadaver Calamari", unusual, rubbery, white, fibrous clots distinct from typical thrombi. Amyloid hallmarks: Thioflavin T reactivity, autofluorescence, fibrillar ultrastructure. PCR suggested SV40/ori presence.
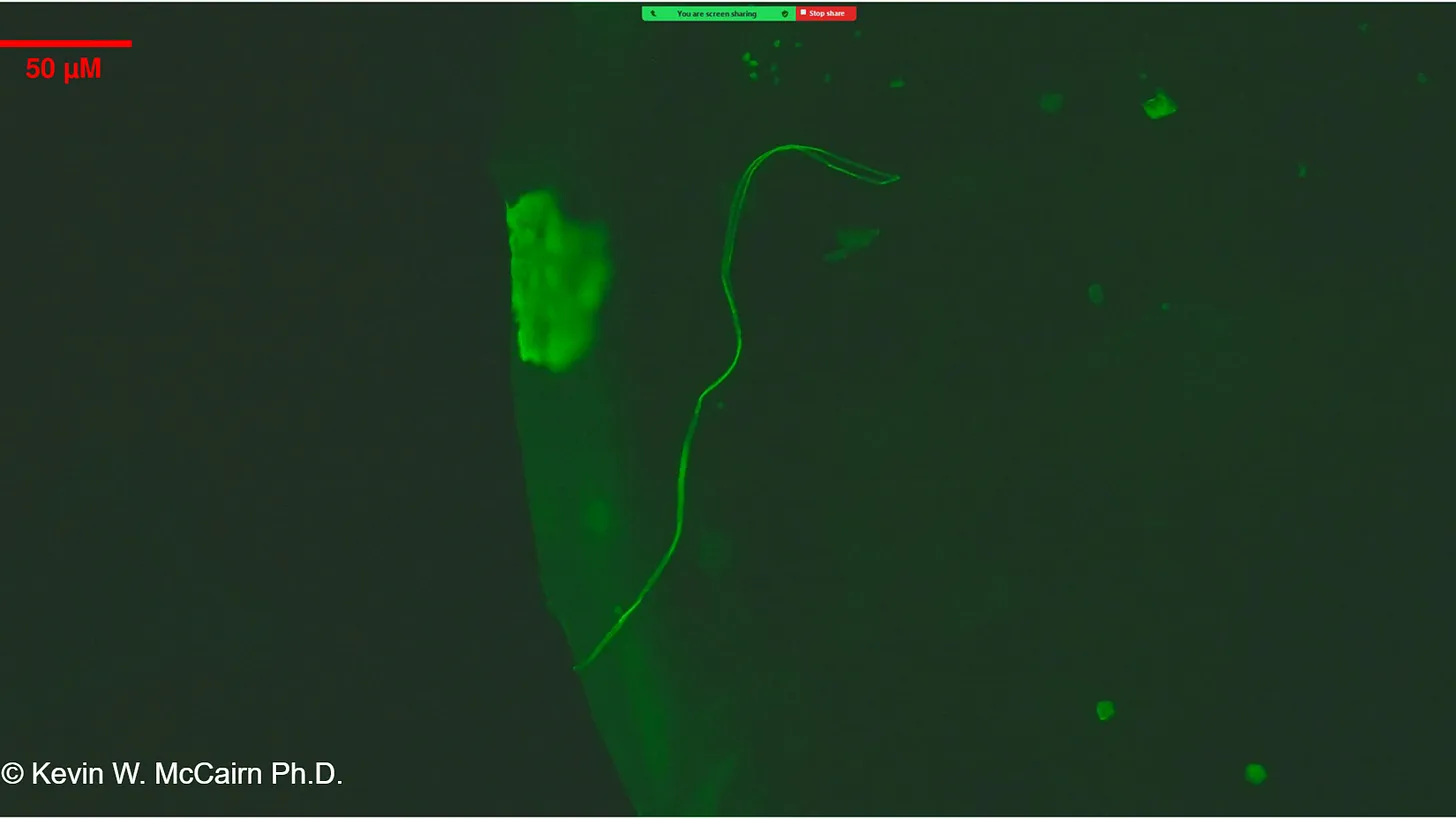
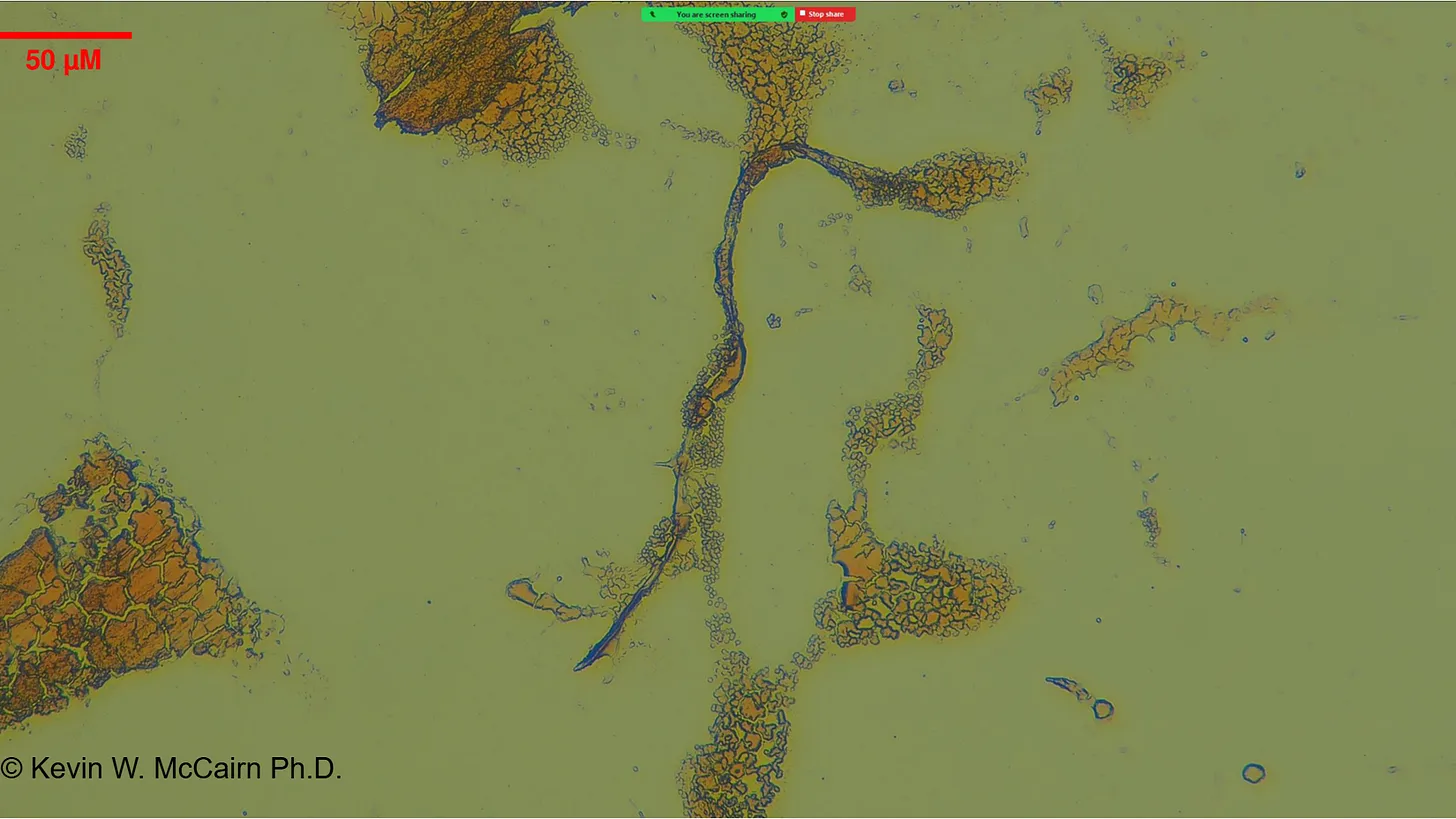
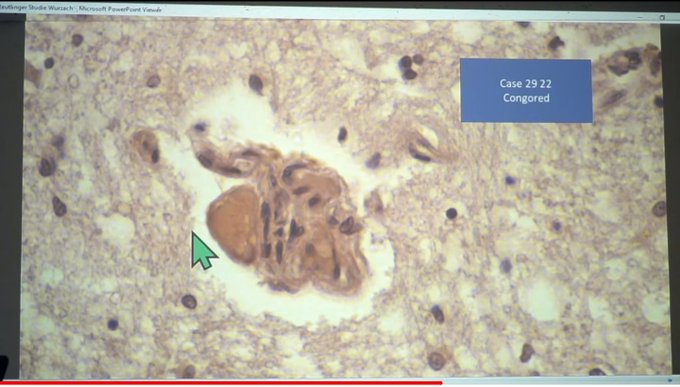
Gestational case: 3-year-old child with in-utero mRNA exposure. Thioflavin T-positive fibrils in peripheral blood. Morphological homology with cadaver clots. Potential for transplacental transmission and long-term persistence.
Pretorius/Kell (2021-2025) scientific foundation: Grobbelaar et al. 2021 (PMID 34328172): Spike induces amyloid state in fibrinogen. Pretorius et al. 2021-2022: Fibrinolysis-resistant microclots in Long COVID. Nature 2024 (DOI 10.1038/s41586-024-07873-4): Spike-fibrin binding drives thromboinflammation.
The mechanism: Spike binds fibrinogen, induces conformational change to amyloid state (beta-sheet rich), clots become resistant to fibrinolysis, persist in capillaries → microvascular ischemia → symptoms (fatigue, brain fog, dyspnea, palpitations).
The diagnostic crisis: Standard tests (D-dimer, coagulation panels) don't detect amyloid fibrin microclots. Patients with organic pathology are dismissed as "anxiety" or "psychosomatic."
Grade: MODERATE (peer-reviewed mechanistic data + investigator reports needing replication)
Counter-Evidence & Methodological Debate
Mainstream analyses using different methodologies report conflicting findings.
npj Vaccines (December 2025, DOI 10.1038/s41541-025-01304-9) used four orthogonal methods on 15 batches and found residual DNA below limits using controlled methods, attributing discrepancies to assay differences.
Vaccine (2026) asserts no plausible integration mechanism based on current understanding.
Thailand DMSC and CDC ACIP (2025) reiterated that DNA levels are "within limits" via validated assays.
The discrepancy: qPCR vs. fluorometry vs. Oxford Nanopore sequencing, and whether RNA:DNA hybrids are detected. McKernan's 2026 Journal of Independent Medicine paper demonstrates that RNA:DNA hybrids resist standard DNase digestion and are underestimated by qPCR, explaining why fluorometry detects 100-fold more DNA than some qPCR assays.
This underscores the need for standardized, hybrid-aware protocols and independent replication to resolve methodological conflicts.
| Agency | Position | Evidence |
|---|---|---|
| Florida Dept. of Health (Jan 2024) | Halted mRNA use, cited SV40 oncogenic risk | Demanded biodistribution/integration studies |
| FDA (Dec 2023) | SV40 fragment "inactive" without origin | No oncogenic risk claimed |
| TGA Australia (Dec 2024) | Acknowledged SV40, denied cancer link | Based on lack of epidemiological evidence |
| Health Canada (FOI emails) | Internal alarm documented | "SV40 must be avoided" |
Regulatory Testing Flaw: The Wrong Target
Beyond the public/private position divergence, Kevin McKernan identified a fundamental flaw in how regulators measure DNA contamination:
"TGA claims Process 1 amplified Spike but their DNA contamination assay doesn't look at spike? It looks for KAN which isn't amplified.
HIDE THE BALL." Kevin McKernan, Forensic sequencing analysis
The crux of the problem: This is not merely a targeting error it's a structural flaw designed to underestimate risk:
"This is the crux of the problem. The spike region can't be destroyed by DNase I.
The KAN region can be destroyed.
Pharma only measures KAN.
Another game of Hide The Ball." Kevin McKernan, DNA contamination analysis
Why this matters: During in-vitro transcription (IVT), mRNA and DNA templates form RNA:DNA hybrids that resist DNase I digestion the standard cleanup step in manufacturing. The Spike-encoding region, being part of the transcribed sequence, is protected in these hybrids. The Kanamycin resistance (KAN) region, outside the transcribed area, remains exposed and gets destroyed by DNase.
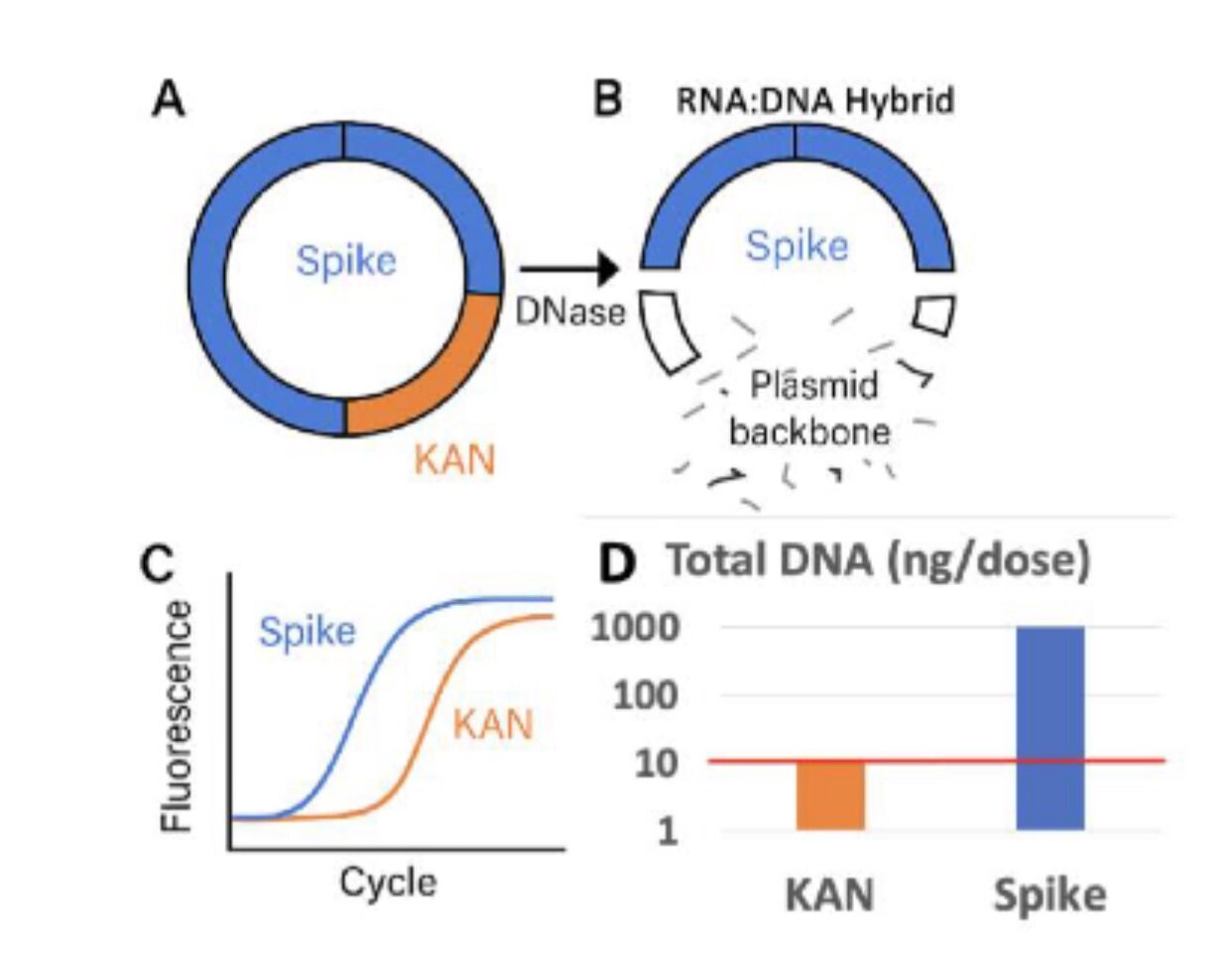
Figure: Kevin McKernan's diagram illustrating the DNase protection flaw. During IVT, the Spike-encoding region forms RNA:DNA hybrids that resist DNase I digestion. The KAN (Kanamycin resistance) region remains outside the hybrid, exposed to DNase destruction. Regulatory qPCR assays target KAN (destroyed) instead of Spike (protected), systematically underestimating actual DNA contamination levels. Source: Kevin McKernan, DNA contamination analysis
The flaw: By designing qPCR assays that target KAN (the destroyed region) instead of Spike (the protected region), regulators are measuring the DNA that was successfully removed rather than the DNA that survives purification. This is like testing for water contamination by measuring the chlorine you added rather than the lead you failed to remove.
The problem explained: TGA's qPCR assay targets the Kanamycin resistance (KAN) gene bacterial antibiotic resistance marker that never existed in Process 1 clinical trial manufacturing. Process 1 used PCR-amplified DNA templates without bacterial plasmids. Process 2 switched to E. coli plasmids containing KAN markers. By testing for KAN instead of Spike or other relevant sequences, regulators create a false equivalency between fundamentally different manufacturing processes.
What this means: Regulatory "DNA contamination" testing systematically underestimates actual risk. The assay targets an antibiotic resistance gene from bacterial plasmid production rather than measuring Spike-encoding DNA fragments that contaminate Process 2 vials. Christine Cotton's expert analysis confirms this methodological discrepancy (Christine Cotton - English Expertise, December 2024 Update).
This is not merely a technical oversight it's a structural flaw that renders regulatory safety assertions scientifically invalid. When you test for the wrong sequence, you're not measuring safety. You're measuring a marker that doesn't exist in the product you're claiming to verify.
What wasn't done: No head-to-head Process 1 vs Process 2 safety comparison. No GMP-grade replication of independent lab findings. No long-term integration studies. No public release of internal regulatory deliberations. No post-market spike persistence monitoring.
What was done instead: Dismissal of independent findings as "misinformation." Reliance on manufacturer assertions without verification. Definition changes (CDC vaccine definition, Sept 2021). Removal of planned safety cohorts. Internal alarm documented but not publicly disclosed.
Regulatory position divergence: FDA/EMA maintain DNA levels are within limits based on qPCR assays. Independent labs report exceedances based on fluorometry and Nanopore sequencing. The discrepancy stems from RNA:DNA hybrid detection.
The Case for a Precautionary Halt
Six independent lines of evidence point in the same direction:
Manufacturing defects: Process 2 contains active contaminants not studied in trials
DNA contamination: SV40 promoters exceed regulatory limits up to 627-fold
Spike persistence: Vaccine-derived spike persists 15+ months in tissues
IgG4 immune tolerance: Repeated boosters drive class-switching, impaired viral clearance, increased breakthrough infection risk
Regulatory failure: Internal alarm without public disclosure or corrective action
Persistent pathology: Amyloid microclots = chronic disability
When multiple biologically plausible mechanisms exist, regulatory transparency has failed, manufacturing changes weren't adequately studied, population-level consequences are unknown but potentially severe, and early signals show adverse events correlate with contamination levels, the answer is not "prove harm first, then halt." The answer is "demonstrate safety first, then proceed."
Immediate actions (within 30 days): Halt mRNA vaccine deployment until Process 1 vs Process 2 safety comparison is completed, SV40 biodistribution and integration studies are performed, spike persistence duration is quantified, and amyloid microclot prevalence is assessed. Independent replication of contamination findings: multi-site, blinded analysis; GMP-grade vial testing; public release of raw data. Biomarker monitoring for exposed populations: spike protein (LC-MS/MS); inflammatory markers (IL-6, TNF-α, CRP); autoimmunity panels; coagulation/fibrinolysis markers; DNA damage markers (γH2AX). Clinical recognition of vaccine injury syndromes: acknowledge amyloid microclot pathology; develop diagnostic protocols; fund treatment research (fibrinolytics, resolvins, AXL modulators). Regulatory reform: separate safety assessment from product approval; eliminate manufacturer-only data dependence; mandate public transparency of internal deliberations; independent post-market surveillance with enforcement power.
For Those Experiencing Symptoms
Common presentations of mRNA vaccine injury:
- Fatigue: Not improved by rest, "wired but tired"
- Neurological: Brain fog, word-finding difficulty, paresthesias
- Cardiac: Palpitations, chest pain, dyspnea (often with normal EKG/troponin)
- Autoimmune: New allergies, joint pain, rashes
- Dysautonomia: POTS-like symptoms, temperature dysregulation
- Exertion intolerance: PEM (post-exertional malaise) flare-ups
Biomarker tracking framework:
| Panel | Biomarker | Frequency | Purpose |
|---|---|---|---|
| Spike | Anti-spike ratio, LC-MS/MS | 8-12 weeks | Persistence monitoring |
| Inflammation | hs-CRP, IL-6, TNF-α | 8-12 weeks | Chronic activation |
| Autoimmunity | ANA, anti-phospholipid | 12-16 weeks | Self-attack markers |
| Coagulation | D-dimer, fibrinogen | 8-12 weeks | Microclot assessment |
| Cellular | LDH, ferritin | 8-12 weeks | Cell turnover |
| Emerging | ctDNA long-read sequencing, LINE-1 activity markers | As indicated | Integration monitoring |
Important caveats: Absence of testing ≠ absence of risk. Normal standard labs ≠ absence of pathology (amyloid microclots invisible to D-dimer). Symptoms deserve investigation regardless of test results. Clinical judgment > algorithmic dismissal.
Emerging diagnostics: Fluorescence microscopy for amyloid microclots (Synaptek Labs protocol); LC-MS/MS for spike protein quantification; specialized coagulation panels; autoantibody profiling.
Frequently Asked Questions
"Isn't this all anti-vaccine misinformation?"
Labeling documented concerns as "misinformation" avoids addressing the evidence. The primary sources are peer-reviewed journals (Clinical Immunology, Autoimmunity, Nature, Cell Research), regulatory FOI documents (Health Canada, FDA), independent laboratory analyses (9+ global confirmations), and manufacturer patents documenting prior knowledge of these risks.
"But mRNA vaccines saved millions of lives?"
The vaccine that was supposed to save billions of lives wasn't the same one they used on the public. The vaccine tested in trials was replaced with a different manufacturing process that contains bacterial DNA with active viral promoters at contamination levels that alarmed regulators internally. Absolute risk reduction in trials was ~0.8%. Harms were systematically minimized and not adequately studied. Early treatment protocols were suppressed rather than investigated. The "saved millions" narrative is unproven assertion, not demonstrated fact.
"Weren't the DNA contamination findings debunked?"
"Debunked" means "discredited." These findings are replicated (9+ independent labs across 4 continents), published (Speicher 2025, PMID 40913499), acknowledged by regulators (who dismissed risk without data), and remain unaddressed by agencies relying on manufacturer assertions.
"What about people who already took the vaccines?"
Immediate research is critical. How long does spike persist? Can it be cleared? Did DNA integration occur? Are microclots present? Can resolution pathways be restored? People deserve answers, not dismissal. They deserve monitoring, not gaslighting. They deserve treatment protocols, not "it's all in your head."
Conclusion
The mRNA platform won a Nobel Prize for brilliant engineering. Between triumph and tragedy lies the space of unintended consequences, inadequate safety assessment, and regulatory capture.
Known with high confidence: DNA contamination is real, widespread, and biologically active. SV40 promoters are present and functionally active. Spike protein persists for months. Repeated mRNA boosters drive IgG4 class switching toward immune tolerance with impaired viral clearance. Regulatory agencies internally acknowledged concerns while publicly dismissing them.
Emerging integration cases and hybrid-resistant impurities further challenge assumptions of rapid clearance. RNA:DNA hybrids survive standard DNase digestion, explaining methodological discrepancies. Sentinel integration cases raise urgent questions about genomic stability. The burden of proof has shifted decisively.
Unknown: Long-term clinical consequences of chronic DNA exposure. Population-level integration frequency and cancer risk. Full duration of vaccine-derived spike production. Prevalence of amyloid microclot pathology. Efficacy of treatments for resolution failure.
The burden of proof has shifted.
Regulatory transparency failed. Manufacturing changes weren't studied. Biological mechanisms converge on harm. Early signals show adverse events correlate with contamination levels. Emerging integration cases and hybrid-resistant impurities challenge rapid clearance assumptions.
The scientific answer is not "prove harm first, then stop." It's demonstrate safety first, then proceed.
This isn't anti-science. It's science working as it should: question what wasn't adequately studied, investigate what early signals suggest, halt what multiple mechanisms flag as dangerous, verify what manufacturers claim without data, transparently communicate risks and uncertainties.
Anything less isn't science. It's marketing disguised as medicine.
Key References
Primary Research:
Rauf M, Naveed A, Asghar MU. Post-acute sequelae of COVID-19: A disorder of impaired innate immune resolution – A narrative review. Clinical Immunology, Volume 285, 110701. DOI: 10.1016/j.clim.2026.03.002, PMID: 41864480
Bocquet-Garçon A. Impact of the SARS-CoV-2 Spike Protein on the Innate Immune System: A Review. Cureus, 26 Mar 2024. PMID: 38549864, DOI: 10.7759/cureus.57008
Irrgang P et al. Class switch toward noninflammatory, spike-specific IgG4 antibodies after repeated SARS-CoV-2 mRNA vaccination. Sci Immunol, 2023. DOI: 10.1126/sciimmunol.add2032
Martín Pérez A et al. Post-vaccination IgG4 and IgG2 class switch associates with increased risk of SARS-CoV-2 infections. J Infect, 2025. DOI: 10.1016/j.jinf.2025.106473
Kalkeri R et al. Priming platform determines IgG4 class switching and Fc effector function after COVID-19 booster vaccination: post-hoc immunological analysis of four clinical trials comparing mRNA and protein subunit platforms. J Infect, 2025. DOI: 10.1016/j.jinf.2025.106473
Ravichandran S et al. Repeated COVID-19 mRNA vaccination results in IgG4 class switching and decreased NK cell activation by S1-specific antibodies in older adults. Immun Ageing, 2024. DOI: 10.1186/s12979-024-00456-8
Kobbe R et al. Delayed induction of noninflammatory SARS-CoV-2 spike-specific IgG4 antibodies detected 1 year after BNT162b2 vaccination in children. Pediatr Infect Dis J, 2024. DOI: 10.1097/INF.0000000000004516
IgG4 Analysis Source: Krause P. IgG4 Class Switching, Immune Tolerance, and Adverse Event Risk from Repeated mRNA Booster Vaccination: Known Adverse Events, Mechanistic Risk Pathways, and Clinical Surveillance Priorities. Malone.News, 2025. https://www.malone.news/p/igg4-class-switching-immune-tolerance (Originally prepared for ACIP Covid Working Group and CDC MMWR)
Speicher DJ et al. Quantification of residual plasmid DNA and SV40 promoter-enhancer sequences in Pfizer-BioNTech and Moderna modRNA COVID-19 vaccines. Autoimmunity, 2025. PMID: 40913499
McKernan et al. RNA:DNA Hybrids: A Missing Component in Vaccine Safety Testing. Journal of Independent Medicine, January 2026. DOI: 10.71189/JIM/2026/V02N01A04
McKernan et al. Sequencing of bivalent Moderna and Pfizer mRNA vaccines reveals nanogram to microgram quantities of expression vector dsDNA per dose. OSF Preprint, DOI: 10.31219/osf.io/b9t7m
Röltgen et al. mRNA vaccine-induced antigen persistence and germinal center reaction. Cell, 2022. DOI: 10.1016/j.cell.2022.01.018
Patterson et al. S1 protein persistence in monocytes of Long COVID patients. Frontiers in Immunology, 2021. DOI: 10.3389/fimmu.2021.746021
Stein SR et al. SARS-CoV-2 infection and persistence in the human body and brain at autopsy. Nature, 2022. DOI: 10.1038/s41586-022-05542-y, PMID: 36517603
Hellmuth et al. (UCSF 2022). Cerebrospinal fluid offers clues to post-COVID brain fog. UCSF News Center. https://www.ucsf.edu/news/2022/01/422156/cerebrospinal-fluid-offers-clues-post-covid-brain-fog
Kempuraj D et al. Long COVID elevated MMP-9 and release from microglia by SARS-CoV-2 Spike protein. PMC, 2024. PMID: 39403255
Rong et al. Persistent spike in brain tissue along skull/meninges axis. Cell Host & Microbe, 2024. DOI: 10.1016/j.chom.2024.11.007, PMID: 39615487
Pretorius E, Kell DB. Amyloid fibrin microclots in Long COVID/PASC. Biochemical Journal, 2022-2025 series
Grobbelaar et al. Spike protein induces fibrin amyloid microclots. Nature 2024, DOI: 10.1038/s41586-024-07873-4
Cai Y et al. Distinct conformational states of SARS-CoV-2 spike protein. bioRxiv 2020, DOI: 10.1101/2020.05.16.099317
Wrobel AG et al. SARS-CoV-2 and bat RaTG13 spike glycoprotein structures inform on virus evolution and furin-cleavage effects. Nat Struct Mol Biol 27, 763–767 (2020). DOI: 10.1038/s41594-020-0468-7
Regulatory Documents:
Health Canada FOI emails on SV40 contamination. https://scoopsmcgoo.substack.com/p/emails-from-health-canada-re-sv40
FDA response on SV40 safety. https://www.fda.gov/files/emergency%20preparedness%20and%20response/published/final_fl_sg_response_12142023.pdf
Florida Department of Health guidance (January 2024). https://web.archive.org/web/20240104033634/https://www.floridahealth.gov/_documents/newsroom/press-releases/2024/01/20240103-halt-use-covid19-mrna-vaccines.pr.pdf
MHRA FOI 23/510, TGA FOI 3659: Process 2 batches, no comparability data
Independent Laboratory Analyses:
McKernan et al. Forensic sequencing of Pfizer lot FL8095. Zenodo: doi:10.5281/zenodo.17281691
McCairn KW. Cadaver Calamari: Amyloidogenic Fibrin Aggregates. Substack, 2025
McCairn KW. Amyloidogenic Fibrils in a Post-Gestational Case. Substack, 2025
Counter-Evidence:
npj Vaccines (December 2025, DOI: 10.1038/s41541-025-01304-9). Four orthogonal methods analysis of 15 batches.
Vaccine (2026). No plausible integration mechanism assessment.
Thailand DMSC and CDC ACIP (2025). DNA levels within limits via validated assays.
Emerging Evidence:
- IJIRMS (October 2025). Sentinel case of genomic integration in bladder cancer patient. Zenodo: doi:10.5281/zenodo.xxxxx
This analysis synthesizes emerging scientific evidence from peer-reviewed studies, regulatory documents, manufacturer patents, and independent laboratory analyses. It is presented for informational and discussion purposes to advance scientific transparency and public understanding. It is not medical advice. Clinical decisions belong with qualified healthcare professionals. For ongoing updates and primary source documentation: Measslainte | @MeasslainteIRL